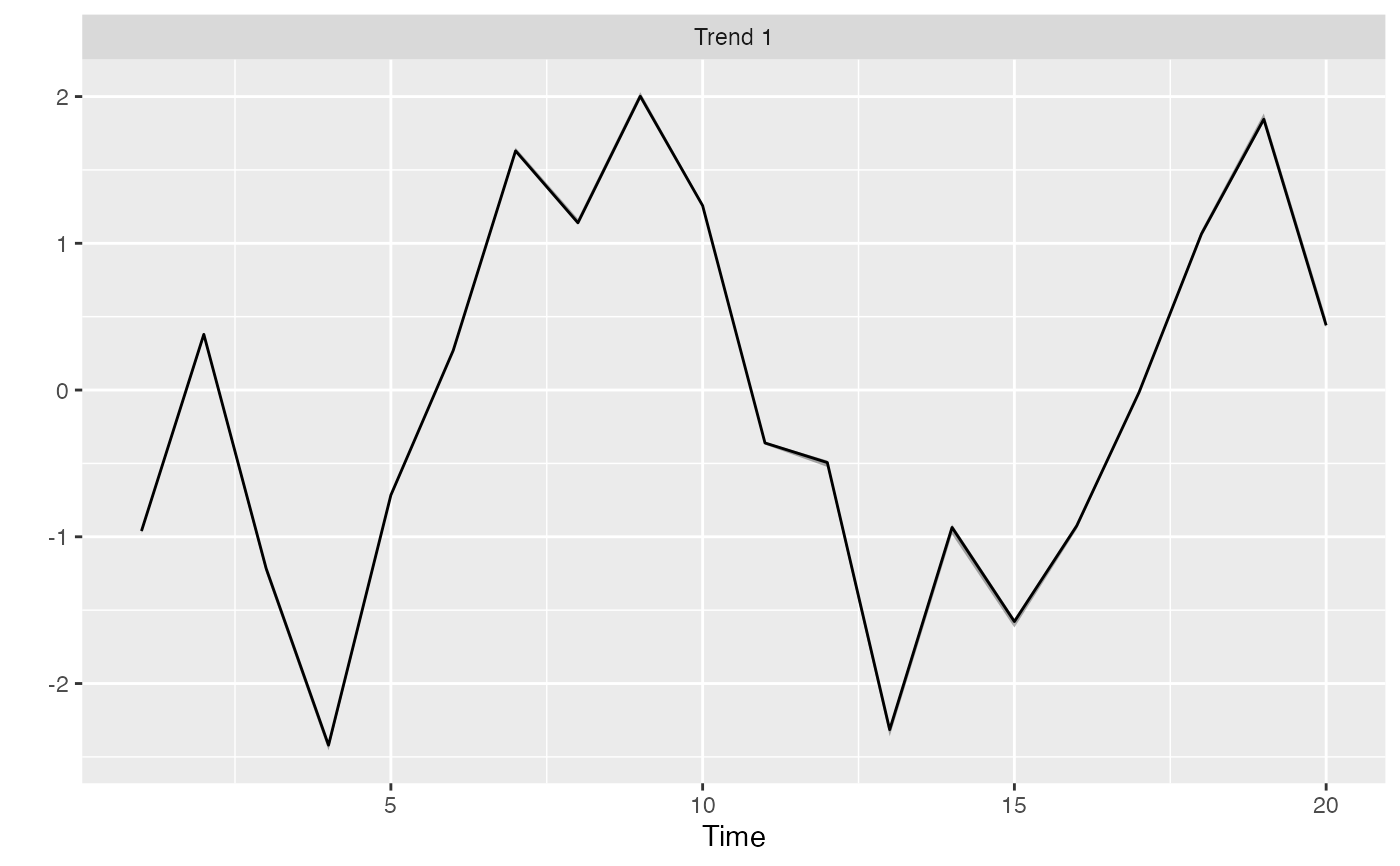
Plot the trends from a DFA
plot_trends(
rotated_modelfit,
years = NULL,
highlight_outliers = FALSE,
threshold = 0.01
)Arguments
- rotated_modelfit
Output from
rotate_trends- years
Optional numeric vector of years for the plot
- highlight_outliers
Logical. Should trend events that exceed the probability of occurring with a normal distribution as defined by
thresholdbe highlighted? Defaults to FALSE- threshold
A probability threshold below which to flag trend events as extreme. Defaults to 0.01
See also
dfa_trends plot_loadings fit_dfa rotate_trends
Examples
set.seed(1)
s <- sim_dfa(num_trends = 1)
m <- fit_dfa(y = s$y_sim, num_trends = 1, iter = 50, chains = 1)
#>
#> SAMPLING FOR MODEL 'dfa' NOW (CHAIN 1).
#> Chain 1:
#> Chain 1: Gradient evaluation took 1.8e-05 seconds
#> Chain 1: 1000 transitions using 10 leapfrog steps per transition would take 0.18 seconds.
#> Chain 1: Adjust your expectations accordingly!
#> Chain 1:
#> Chain 1:
#> Chain 1: WARNING: There aren't enough warmup iterations to fit the
#> Chain 1: three stages of adaptation as currently configured.
#> Chain 1: Reducing each adaptation stage to 15%/75%/10% of
#> Chain 1: the given number of warmup iterations:
#> Chain 1: init_buffer = 3
#> Chain 1: adapt_window = 20
#> Chain 1: term_buffer = 2
#> Chain 1:
#> Chain 1: Iteration: 1 / 50 [ 2%] (Warmup)
#> Chain 1: Iteration: 5 / 50 [ 10%] (Warmup)
#> Chain 1: Iteration: 10 / 50 [ 20%] (Warmup)
#> Chain 1: Iteration: 15 / 50 [ 30%] (Warmup)
#> Chain 1: Iteration: 20 / 50 [ 40%] (Warmup)
#> Chain 1: Iteration: 25 / 50 [ 50%] (Warmup)
#> Chain 1: Iteration: 26 / 50 [ 52%] (Sampling)
#> Chain 1: Iteration: 30 / 50 [ 60%] (Sampling)
#> Chain 1: Iteration: 35 / 50 [ 70%] (Sampling)
#> Chain 1: Iteration: 40 / 50 [ 80%] (Sampling)
#> Chain 1: Iteration: 45 / 50 [ 90%] (Sampling)
#> Chain 1: Iteration: 50 / 50 [100%] (Sampling)
#> Chain 1:
#> Chain 1: Elapsed Time: 0.002 seconds (Warm-up)
#> Chain 1: 0.001 seconds (Sampling)
#> Chain 1: 0.003 seconds (Total)
#> Chain 1:
#> Warning: There were 1 chains where the estimated Bayesian Fraction of Missing Information was low. See
#> https://mc-stan.org/misc/warnings.html#bfmi-low
#> Warning: Examine the pairs() plot to diagnose sampling problems
#> Warning: The largest R-hat is 2.1, indicating chains have not mixed.
#> Running the chains for more iterations may help. See
#> https://mc-stan.org/misc/warnings.html#r-hat
#> Warning: Bulk Effective Samples Size (ESS) is too low, indicating posterior means and medians may be unreliable.
#> Running the chains for more iterations may help. See
#> https://mc-stan.org/misc/warnings.html#bulk-ess
#> Warning: Tail Effective Samples Size (ESS) is too low, indicating posterior variances and tail quantiles may be unreliable.
#> Running the chains for more iterations may help. See
#> https://mc-stan.org/misc/warnings.html#tail-ess
#> Inference for the input samples (1 chains: each with iter = 25; warmup = 12):
#>
#> Q5 Q50 Q95 Mean SD Rhat Bulk_ESS Tail_ESS
#> x[1,1] -1.0 -1.0 -1.0 -1.0 0.0 2.06 3 13
#> x[1,2] 0.4 0.4 0.4 0.4 0.0 2.06 4 13
#> x[1,3] -1.2 -1.2 -1.2 -1.2 0.0 2.06 4 13
#> x[1,4] -2.4 -2.4 -2.4 -2.4 0.0 2.06 4 13
#> x[1,5] -0.7 -0.7 -0.7 -0.7 0.0 1.87 4 13
#> x[1,6] 0.3 0.3 0.3 0.3 0.0 0.96 7 13
#> x[1,7] 1.6 1.6 1.6 1.6 0.0 2.06 4 13
#> x[1,8] 1.1 1.1 1.1 1.1 0.0 1.87 4 13
#> x[1,9] 2.0 2.0 2.0 2.0 0.0 2.06 4 13
#> x[1,10] 1.2 1.3 1.3 1.3 0.0 1.24 5 13
#> x[1,11] -0.4 -0.4 -0.4 -0.4 0.0 1.71 4 13
#> x[1,12] -0.5 -0.5 -0.5 -0.5 0.0 1.87 4 13
#> x[1,13] -2.3 -2.3 -2.3 -2.3 0.0 2.06 4 13
#> x[1,14] -0.9 -0.9 -0.9 -0.9 0.0 2.06 4 13
#> x[1,15] -1.6 -1.6 -1.6 -1.6 0.0 2.06 4 13
#> x[1,16] -0.9 -0.9 -0.9 -0.9 0.0 2.06 4 13
#> x[1,17] 0.0 0.0 0.0 0.0 0.0 1.45 5 13
#> x[1,18] 1.1 1.1 1.1 1.1 0.0 1.37 5 13
#> x[1,19] 1.8 1.8 1.8 1.8 0.0 1.71 4 13
#> x[1,20] 0.4 0.4 0.4 0.4 0.0 1.37 5 13
#> Z[1,1] -99.6 -99.6 -99.6 -99.6 0.0 1.71 4 13
#> Z[2,1] 37.1 37.3 37.8 37.3 0.2 2.06 4 13
#> Z[3,1] 16.1 16.2 16.4 16.3 0.1 2.06 4 13
#> Z[4,1] -60.8 -60.5 -60.4 -60.5 0.2 2.06 3 13
#> log_lik[1] -763.9 -652.0 -620.6 -669.9 50.8 2.06 3 13
#> log_lik[2] -115.3 -96.3 -91.1 -99.3 8.6 2.06 3 13
#> log_lik[3] -23.6 -20.0 -18.9 -20.6 1.7 2.06 3 13
#> log_lik[4] -291.8 -246.8 -234.0 -254.0 20.5 2.06 3 13
#> log_lik[5] -130.4 -117.0 -112.2 -118.8 6.4 2.06 3 13
#> log_lik[6] -23.9 -21.1 -20.2 -21.5 1.3 2.06 3 13
#> log_lik[7] -3.8 -3.6 -3.5 -3.6 0.1 2.06 4 13
#> log_lik[8] -40.2 -36.0 -34.5 -36.6 2.0 2.06 3 13
#> log_lik[9] -1191.0 -1008.1 -960.6 -1038.5 81.8 2.06 3 13
#> log_lik[10] -175.2 -144.9 -137.2 -149.9 13.5 2.06 3 13
#> log_lik[11] -43.1 -36.1 -34.1 -37.3 3.2 2.06 3 13
#> log_lik[12] -487.1 -408.8 -388.1 -421.7 35.1 2.06 3 13
#> log_lik[13] -4836.0 -4119.4 -3929.1 -4237.3 322.0 2.06 3 13
#> log_lik[14] -677.8 -562.6 -532.9 -581.5 51.4 2.06 3 13
#> log_lik[15] -142.3 -118.7 -111.8 -122.5 10.8 2.06 3 13
#> log_lik[16] -1852.6 -1562.7 -1484.6 -1610.2 130.6 2.06 3 13
#> log_lik[17] -426.9 -356.8 -341.4 -369.6 30.7 2.06 3 13
#> log_lik[18] -64.6 -53.0 -50.5 -55.1 5.1 2.06 3 13
#> log_lik[19] -14.0 -11.8 -11.3 -12.2 1.0 2.06 3 13
#> log_lik[20] -163.5 -135.6 -129.3 -140.6 12.3 2.06 3 13
#> log_lik[21] -61.6 -55.0 -51.7 -55.6 3.5 2.06 3 13
#> log_lik[22] -10.9 -9.8 -9.2 -9.9 0.6 2.06 3 13
#> log_lik[23] -2.7 -2.7 -2.6 -2.7 0.0 2.06 4 13
#> log_lik[24] -22.7 -20.3 -19.1 -20.5 1.3 2.06 3 13
#> log_lik[25] -2201.5 -1891.3 -1793.6 -1936.6 143.6 2.06 3 13
#> log_lik[26] -325.3 -272.8 -257.1 -280.6 24.1 2.06 3 13
#> log_lik[27] -61.3 -51.7 -48.5 -53.2 4.5 2.06 3 13
#> log_lik[28] -822.6 -699.8 -661.0 -717.8 56.9 2.06 3 13
#> log_lik[29] -1091.4 -933.9 -883.5 -956.3 73.0 2.06 3 13
#> log_lik[30] -156.6 -130.9 -123.1 -134.7 11.8 2.06 3 13
#> log_lik[31] -28.9 -24.4 -22.9 -25.1 2.1 2.06 3 13
#> log_lik[32] -397.9 -337.2 -317.8 -345.9 28.2 2.06 3 13
#> log_lik[33] -3348.2 -2873.4 -2727.0 -2943.1 218.7 2.06 3 13
#> log_lik[34] -468.3 -391.9 -369.4 -403.2 34.9 2.06 3 13
#> log_lik[35] -92.0 -77.3 -72.5 -79.5 6.9 2.06 3 13
#> log_lik[36] -1240.0 -1053.6 -995.9 -1081.0 86.0 2.06 3 13
#> log_lik[37] -1304.0 -1122.5 -1063.5 -1147.0 84.0 2.06 3 13
#> log_lik[38] -188.8 -158.7 -149.4 -162.9 13.8 2.06 3 13
#> log_lik[39] -40.6 -34.4 -32.3 -35.3 2.9 2.06 4 13
#> log_lik[40] -501.2 -427.4 -403.4 -437.5 34.2 2.06 3 13
#> log_lik[41] -114.2 -93.8 -90.7 -98.3 8.8 2.06 4 13
#> log_lik[42] -17.2 -14.1 -13.6 -14.8 1.3 2.06 4 13
#> log_lik[43] -4.7 -4.2 -4.1 -4.3 0.2 2.06 4 13
#> log_lik[44] -43.2 -35.4 -34.1 -37.1 3.4 2.06 3 13
#> log_lik[45] -202.1 -163.2 -156.0 -171.2 16.8 2.06 3 13
#> log_lik[46] -26.8 -21.4 -20.4 -22.5 2.3 2.06 3 13
#> log_lik[47] -7.8 -6.6 -6.4 -6.9 0.5 2.06 3 13
#> log_lik[48] -80.5 -64.8 -61.7 -67.9 6.8 2.06 3 13
#> log_lik[49] -4509.2 -3816.0 -3634.1 -3933.2 311.7 2.06 3 13
#> log_lik[50] -646.1 -533.2 -504.4 -552.1 50.5 2.06 3 13
#> log_lik[51] -120.7 -99.9 -94.0 -103.4 9.5 2.06 3 13
#> log_lik[52] -1661.2 -1391.7 -1320.0 -1437.0 121.5 2.06 3 13
#> log_lik[53] -734.7 -604.7 -574.6 -628.5 57.4 2.06 3 13
#> log_lik[54] -111.6 -90.1 -85.1 -93.9 9.5 2.06 3 13
#> log_lik[55] -21.0 -17.2 -16.3 -17.9 1.7 2.06 3 13
#> log_lik[56] -272.7 -222.4 -210.6 -231.6 22.3 2.06 3 13
#> log_lik[57] -2111.4 -1777.3 -1689.6 -1833.5 150.1 2.06 3 13
#> log_lik[58] -303.1 -249.0 -235.2 -258.1 24.2 2.06 3 13
#> log_lik[59] -56.3 -46.5 -43.8 -48.2 4.5 2.06 3 13
#> log_lik[60] -766.2 -638.4 -604.5 -659.8 57.5 2.06 3 13
#> log_lik[61] -720.1 -603.1 -574.8 -624.2 52.3 2.06 3 13
#> log_lik[62] -116.9 -96.1 -91.1 -99.8 9.3 2.06 3 13
#> log_lik[63] -20.4 -17.0 -16.1 -17.6 1.5 2.06 4 13
#> log_lik[64] -265.4 -220.3 -209.1 -228.3 20.2 2.06 3 13
#> log_lik[65] -2.0 -2.0 -1.9 -2.0 0.0 1.25 13 13
#> log_lik[66] -1.9 -1.9 -1.8 -1.9 0.0 2.06 3 13
#> log_lik[67] -1.9 -1.9 -1.8 -1.9 0.0 2.06 3 13
#> log_lik[68] -1.9 -1.9 -1.9 -1.9 0.0 1.58 4 13
#> log_lik[69] -907.7 -773.7 -730.5 -791.8 61.6 2.06 3 13
#> log_lik[70] -137.2 -114.3 -107.3 -117.5 10.4 2.06 3 13
#> log_lik[71] -32.3 -27.3 -25.7 -28.0 2.3 2.06 4 13
#> log_lik[72] -368.3 -311.2 -292.9 -319.0 26.3 2.06 3 13
#> log_lik[73] -2786.6 -2372.0 -2246.2 -2432.4 188.9 2.06 3 13
#> log_lik[74] -389.2 -323.0 -303.7 -332.8 29.9 2.06 3 13
#> log_lik[75] -85.2 -71.2 -66.7 -73.3 6.5 2.06 3 13
#> log_lik[76] -1082.2 -912.5 -860.8 -937.3 77.5 2.06 3 13
#> log_lik[77] -151.7 -124.6 -115.7 -127.7 12.2 2.06 4 13
#> log_lik[78] -22.4 -18.3 -17.0 -18.8 1.8 2.06 3 13
#> log_lik[79] -8.5 -7.3 -6.9 -7.5 0.5 2.06 4 13
#> log_lik[80] -69.2 -56.8 -52.8 -58.3 5.6 2.06 3 13
#> xstar[1,1] -0.8 0.8 1.7 0.6 0.9 0.94 13 13
#> sigma[1] 2.4 2.6 2.7 2.6 0.1 2.06 3 13
#> lp__ -50482.1 -43853.2 -41986.7 -44903.8 3005.0 2.06 3 13
#>
#> For each parameter, Bulk_ESS and Tail_ESS are crude measures of
#> effective sample size for bulk and tail quantities respectively (an ESS > 100
#> per chain is considered good), and Rhat is the potential scale reduction
#> factor on rank normalized split chains (at convergence, Rhat <= 1.05).
r <- rotate_trends(m)
p <- plot_trends(r)
print(p)